[](https://pypi.python.org/pypi/chemcloud)
[](https://pypi.python.org/pypi/chemcloud)
[](https://pypi.python.org/pypi/chemcloud)
[](https://github.com/mtzgroup/chemcloud-client/actions)
[](https://github.com/mtzgroup/chemcloud-client/actions)
[](https://github.com/charliermarsh/ruff)
# chemcloud - A Python Client for ChemCloud
`chemcloud` is a python client for the [ChemCloud Server](https://github.com/mtzgroup/chemcloud-server). The client provides a simple yet powerful interface to perform computational chemistry calculations using nothing but modern Python and an internet connection.
**Documentation**: <https://mtzgroup.github.io/chemcloud-client>
`chemcloud` works in harmony with a suite of other quantum chemistry tools for fast, structured, and interoperable quantum chemistry.
## The QC Suite of Programs
- [qcio](https://github.com/coltonbh/qcio) - Elegant and intuitive data structures for quantum chemistry, featuring seamless Jupyter Notebook visualizations. [Documentation](https://qcio.coltonhicks.com)
- [qcparse](https://github.com/coltonbh/qcparse) - A library for efficient parsing of quantum chemistry data into structured `qcio` objects.
- [qcop](https://github.com/coltonbh/qcop) - A package for operating quantum chemistry programs using `qcio` standardized data structures. Compatible with `TeraChem`, `psi4`, `QChem`, `NWChem`, `ORCA`, `Molpro`, `geomeTRIC`, and many more, featuring seamless Jupyter Notebook visualizations.
- [BigChem](https://github.com/mtzgroup/bigchem) - A distributed application for running quantum chemistry calculations at scale across clusters of computers or the cloud. Bring multi-node scaling to your favorite quantum chemistry program, featuring seamless Jupyter Notebook visualizations.
- `ChemCloud` - A [web application](https://github.com/mtzgroup/chemcloud-server) and associated [Python client](https://github.com/mtzgroup/chemcloud-client) for exposing a BigChem cluster securely over the internet, featuring seamless Jupyter Notebook visualizations.
## Installation
```sh
pip install chemcloud
```
## Quickstart
- Create a ChemCloud account at [https://chemcloud.mtzlab.com/signup](https://chemcloud.mtzlab.com/signup) (or the address of the ChemCloud Server you want to communicate with).
- Instantiate a client
- Configure client (only required the very first time you use `CCClient`)
```python
from chemcloud import CCClient
client = CCClient()
client.configure() # only run this the very first time you use CCClient
# See supported compute engines on the ChemCloud Server
client.supported_engines
['psi4', 'terachem', ...]
# Test connection to ChemCloud
client.hello_world("Colton")
'Welcome to ChemCloud, Colton'
```
- Run calculations just like you would with `qcop` except calling `client.compute` instead of `qcop.compute`. Rather than getting back an `Output` object directly, `client.compute` returns a `FutureOutput` object which can be used to get the output of the computation once it is complete.
```python
from qcio import Structure, ProgramInput
from chemcloud import CCClient
client = CCClient()
# Create the structure
h2o = Structure.open("h2o.xyz")
# Define the program input
prog_input = ProgramInput(
structure=h2o,
calctype="energy",
model={"method": "hf", "basis": "sto-3g"},
keywords={"purify": "no", "restricted": False},
)
# Submit the calculation to the server
future_output = client.compute("terachem", prog_input, collect_files=True)
# Status can be checked at any time
future_result.status
# Get the output (blocking)
output = future_output.get()
# Inspect the output
output.input_data # Input data used by the QC program
output.success # Whether the calculation succeeded
output.results # All structured results from the calculation
output.stdout # Stdout log from the calculation
output.pstdout # Shortcut to print out the stdout in human readable format
output.files # Any files returned by the calculation
output.provenance # Provenance information about the calculation
output.extras # Any extra information not in the schema
output.traceback # Stack trace if calculation failed
output.ptraceback # Shortcut to print out the traceback in human readable format
```
## Examples
Examples of various computations can be found in the [examples directory](https://github.com/mtzgroup/chemcloud-client/tree/main/examples).
## ✨ Visualization ✨
Visualize all your results with a single line of code!
First install the visualization module:
```sh
pip install qcio[view]
```
or if your shell requires `''` around arguments with brackets:
```sh
pip install 'qcio[view]'
```
Then in a Jupyter notebook import the `qcio` view module and call `view.view(...)` passing it one or any number of `qcio` objects you want to visualizing including `Structure` objects or any `ProgramOutput` object. You may also pass an array of `titles` and/or `subtitles` to add additional information to the molecular structure display. If no titles are passed `qcio` with look for `Structure` identifiers such as a name or SMILES to label the `Structure`.
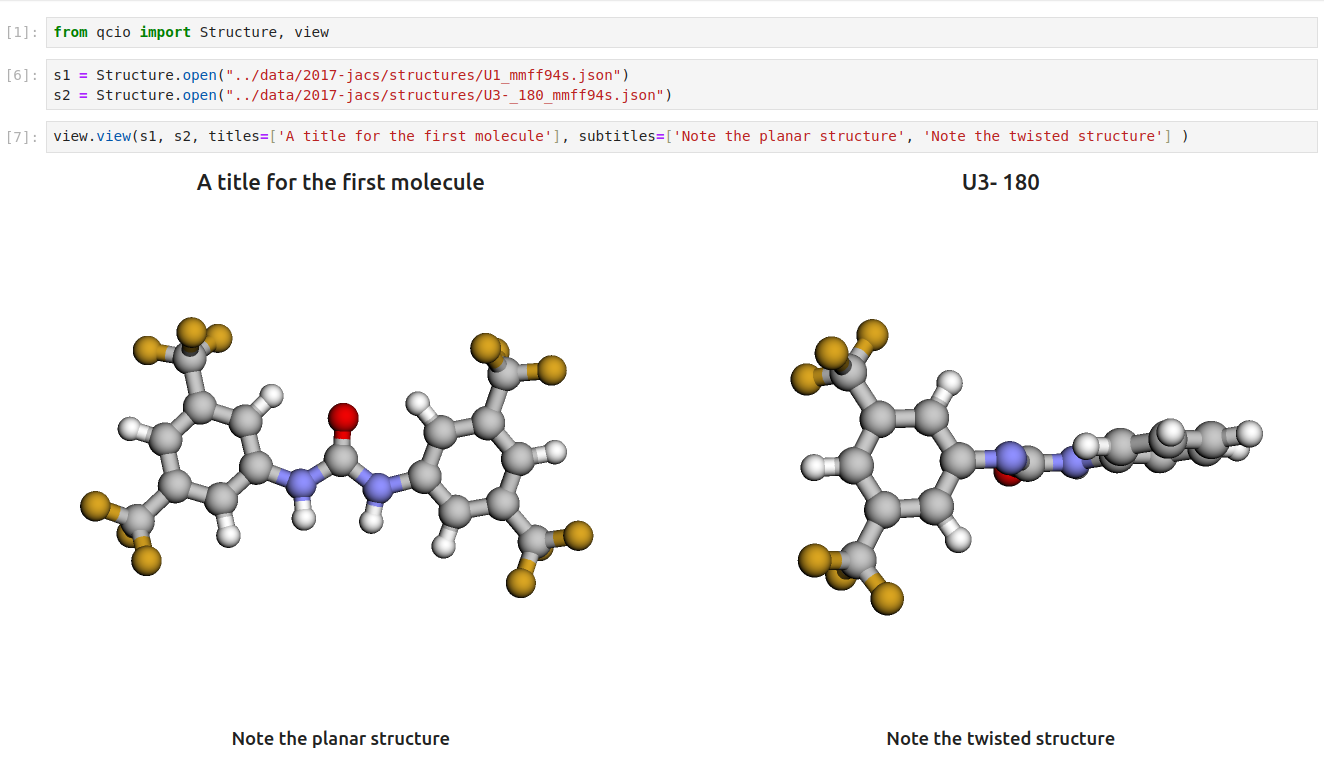
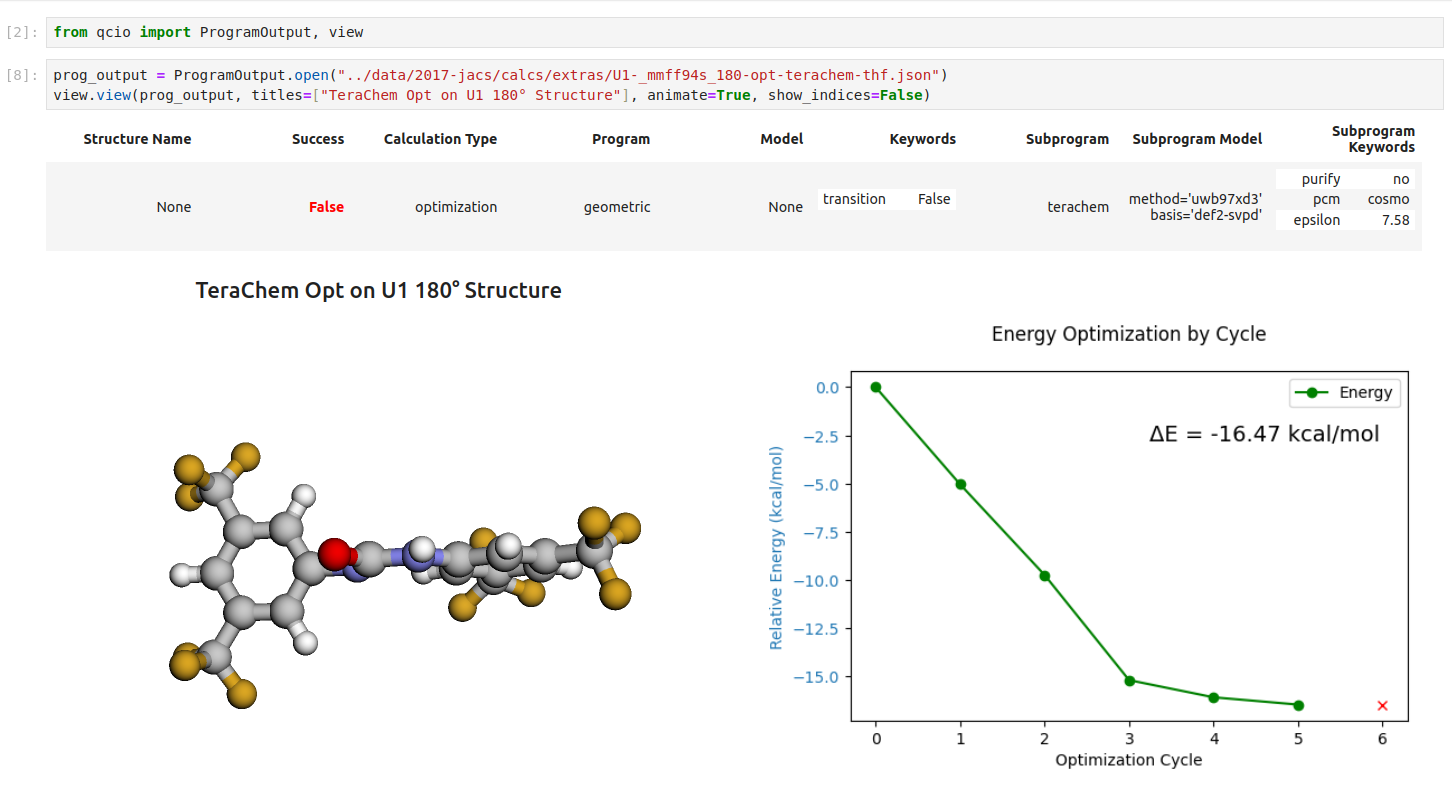
Seamless visualizations for `ProgramOutput` objects make results analysis easy!
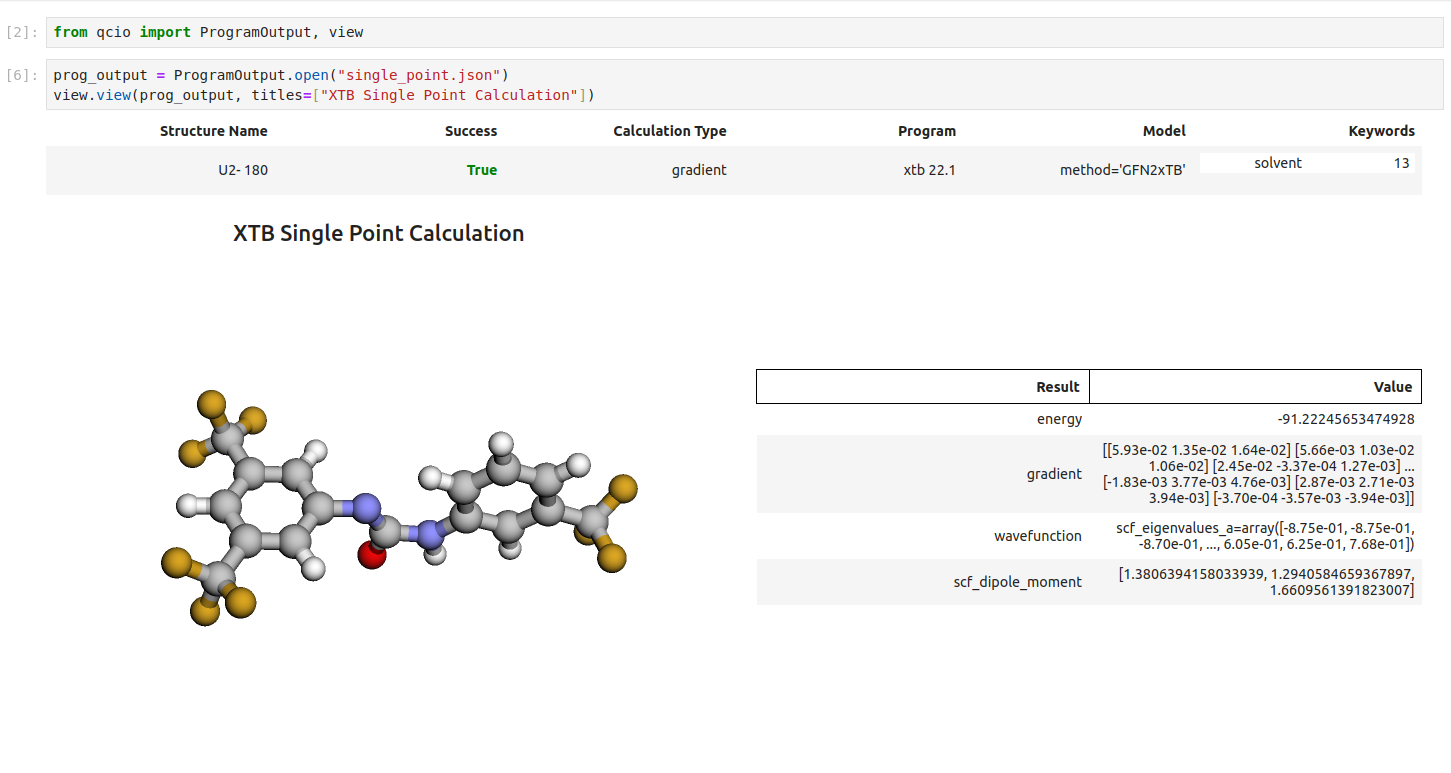
Single point calculations display their results in a table.
If you want to use the HTML generated by the viewer to build your own dashboards use the functions inside of `qcio.view.py` that begin with the word `generate_` to create HTML you can insert into any dashboard.
## Support
If you have any issues with `chemcloud` or would like to request a feature, please open an [issue](https://github.com/mtzgroup/chemcloud-client/issues).
Raw data
{
"_id": null,
"home_page": "https://mtzgroup.github.io/chemcloud-client/",
"name": "chemcloud",
"maintainer": null,
"docs_url": null,
"requires_python": "<4.0,>=3.9",
"maintainer_email": null,
"keywords": null,
"author": "Colton Hicks",
"author_email": "github@coltonhicks.com",
"download_url": "https://files.pythonhosted.org/packages/47/78/41ae01813769ab5910677d6becc81fd93105bafc5851692cd38e3cd35759/chemcloud-0.12.1.tar.gz",
"platform": null,
"description": "[](https://pypi.python.org/pypi/chemcloud)\n[](https://pypi.python.org/pypi/chemcloud)\n[](https://pypi.python.org/pypi/chemcloud)\n[](https://github.com/mtzgroup/chemcloud-client/actions)\n[](https://github.com/mtzgroup/chemcloud-client/actions)\n[](https://github.com/charliermarsh/ruff)\n\n# chemcloud - A Python Client for ChemCloud\n\n`chemcloud` is a python client for the [ChemCloud Server](https://github.com/mtzgroup/chemcloud-server). The client provides a simple yet powerful interface to perform computational chemistry calculations using nothing but modern Python and an internet connection.\n\n**Documentation**: <https://mtzgroup.github.io/chemcloud-client>\n\n`chemcloud` works in harmony with a suite of other quantum chemistry tools for fast, structured, and interoperable quantum chemistry.\n\n## The QC Suite of Programs\n\n- [qcio](https://github.com/coltonbh/qcio) - Elegant and intuitive data structures for quantum chemistry, featuring seamless Jupyter Notebook visualizations. [Documentation](https://qcio.coltonhicks.com)\n- [qcparse](https://github.com/coltonbh/qcparse) - A library for efficient parsing of quantum chemistry data into structured `qcio` objects.\n- [qcop](https://github.com/coltonbh/qcop) - A package for operating quantum chemistry programs using `qcio` standardized data structures. Compatible with `TeraChem`, `psi4`, `QChem`, `NWChem`, `ORCA`, `Molpro`, `geomeTRIC`, and many more, featuring seamless Jupyter Notebook visualizations.\n- [BigChem](https://github.com/mtzgroup/bigchem) - A distributed application for running quantum chemistry calculations at scale across clusters of computers or the cloud. Bring multi-node scaling to your favorite quantum chemistry program, featuring seamless Jupyter Notebook visualizations.\n- `ChemCloud` - A [web application](https://github.com/mtzgroup/chemcloud-server) and associated [Python client](https://github.com/mtzgroup/chemcloud-client) for exposing a BigChem cluster securely over the internet, featuring seamless Jupyter Notebook visualizations.\n\n## Installation\n\n```sh\npip install chemcloud\n```\n\n## Quickstart\n\n- Create a ChemCloud account at [https://chemcloud.mtzlab.com/signup](https://chemcloud.mtzlab.com/signup) (or the address of the ChemCloud Server you want to communicate with).\n- Instantiate a client\n- Configure client (only required the very first time you use `CCClient`)\n\n```python\nfrom chemcloud import CCClient\n\nclient = CCClient()\nclient.configure() # only run this the very first time you use CCClient\n# See supported compute engines on the ChemCloud Server\nclient.supported_engines\n['psi4', 'terachem', ...]\n# Test connection to ChemCloud\nclient.hello_world(\"Colton\")\n'Welcome to ChemCloud, Colton'\n```\n\n- Run calculations just like you would with `qcop` except calling `client.compute` instead of `qcop.compute`. Rather than getting back an `Output` object directly, `client.compute` returns a `FutureOutput` object which can be used to get the output of the computation once it is complete.\n\n```python\nfrom qcio import Structure, ProgramInput\nfrom chemcloud import CCClient\n\nclient = CCClient()\n\n# Create the structure\nh2o = Structure.open(\"h2o.xyz\")\n\n# Define the program input\nprog_input = ProgramInput(\n structure=h2o,\n calctype=\"energy\",\n model={\"method\": \"hf\", \"basis\": \"sto-3g\"},\n keywords={\"purify\": \"no\", \"restricted\": False},\n)\n\n# Submit the calculation to the server\nfuture_output = client.compute(\"terachem\", prog_input, collect_files=True)\n# Status can be checked at any time\nfuture_result.status\n# Get the output (blocking)\noutput = future_output.get()\n\n# Inspect the output\noutput.input_data # Input data used by the QC program\noutput.success # Whether the calculation succeeded\noutput.results # All structured results from the calculation\noutput.stdout # Stdout log from the calculation\noutput.pstdout # Shortcut to print out the stdout in human readable format\noutput.files # Any files returned by the calculation\noutput.provenance # Provenance information about the calculation\noutput.extras # Any extra information not in the schema\noutput.traceback # Stack trace if calculation failed\noutput.ptraceback # Shortcut to print out the traceback in human readable format\n```\n\n## Examples\n\nExamples of various computations can be found in the [examples directory](https://github.com/mtzgroup/chemcloud-client/tree/main/examples).\n\n## \u2728 Visualization \u2728\n\nVisualize all your results with a single line of code!\n\nFirst install the visualization module:\n\n```sh\npip install qcio[view]\n```\n\nor if your shell requires `''` around arguments with brackets:\n\n```sh\npip install 'qcio[view]'\n```\n\nThen in a Jupyter notebook import the `qcio` view module and call `view.view(...)` passing it one or any number of `qcio` objects you want to visualizing including `Structure` objects or any `ProgramOutput` object. You may also pass an array of `titles` and/or `subtitles` to add additional information to the molecular structure display. If no titles are passed `qcio` with look for `Structure` identifiers such as a name or SMILES to label the `Structure`.\n\n\n\nSeamless visualizations for `ProgramOutput` objects make results analysis easy!\n\n\n\nSingle point calculations display their results in a table.\n\n\n\nIf you want to use the HTML generated by the viewer to build your own dashboards use the functions inside of `qcio.view.py` that begin with the word `generate_` to create HTML you can insert into any dashboard.\n\n## Support\n\nIf you have any issues with `chemcloud` or would like to request a feature, please open an [issue](https://github.com/mtzgroup/chemcloud-client/issues).\n\n",
"bugtrack_url": null,
"license": "MIT",
"summary": "Fast, secure, scalable quantum chemistry. A Python client for the the ChemCloud server.",
"version": "0.12.1",
"project_urls": {
"Documentation": "https://mtzgroup.github.io/chemcloud-client/",
"Homepage": "https://mtzgroup.github.io/chemcloud-client/",
"Repository": "https://github.com/mtzgroup/chemcloud-client"
},
"split_keywords": [],
"urls": [
{
"comment_text": "",
"digests": {
"blake2b_256": "6a02720d6ca1a38f51441df060432536e5b1273cf632ec3208c676370b7096a8",
"md5": "3d1e5688bb4db196d19a3fcbcab1db8d",
"sha256": "6279ff9ea1c766e301f078b5d3024ed86f1c0ec078a985001317bdfdba24110b"
},
"downloads": -1,
"filename": "chemcloud-0.12.1-py3-none-any.whl",
"has_sig": false,
"md5_digest": "3d1e5688bb4db196d19a3fcbcab1db8d",
"packagetype": "bdist_wheel",
"python_version": "py3",
"requires_python": "<4.0,>=3.9",
"size": 13720,
"upload_time": "2024-09-12T00:08:46",
"upload_time_iso_8601": "2024-09-12T00:08:46.418929Z",
"url": "https://files.pythonhosted.org/packages/6a/02/720d6ca1a38f51441df060432536e5b1273cf632ec3208c676370b7096a8/chemcloud-0.12.1-py3-none-any.whl",
"yanked": false,
"yanked_reason": null
},
{
"comment_text": "",
"digests": {
"blake2b_256": "477841ae01813769ab5910677d6becc81fd93105bafc5851692cd38e3cd35759",
"md5": "a25ae09ae8cbdc948d137b743c1c95dd",
"sha256": "2a860b42afe2d85cdc5dab9148cb36a2710ea4e40dd1356c078995016333a230"
},
"downloads": -1,
"filename": "chemcloud-0.12.1.tar.gz",
"has_sig": false,
"md5_digest": "a25ae09ae8cbdc948d137b743c1c95dd",
"packagetype": "sdist",
"python_version": "source",
"requires_python": "<4.0,>=3.9",
"size": 13926,
"upload_time": "2024-09-12T00:08:47",
"upload_time_iso_8601": "2024-09-12T00:08:47.683875Z",
"url": "https://files.pythonhosted.org/packages/47/78/41ae01813769ab5910677d6becc81fd93105bafc5851692cd38e3cd35759/chemcloud-0.12.1.tar.gz",
"yanked": false,
"yanked_reason": null
}
],
"upload_time": "2024-09-12 00:08:47",
"github": true,
"gitlab": false,
"bitbucket": false,
"codeberg": false,
"github_user": "mtzgroup",
"github_project": "chemcloud-client",
"travis_ci": false,
"coveralls": false,
"github_actions": true,
"lcname": "chemcloud"
}
